
The DNA Damage Response (DDR) pathway is the core biological system through which cells detect and repair genomic lesions. Targeting DDR pathways and inhibiting DNA repair mechanisms has emerged as a highly promising strategy in cancer therapy, with significant breakthroughs achieved in the development of DDR inhibitors. Antibody-drug conjugates (ADCs)which combine the precise targeting of antibodies with the potent cytotoxicity of drugsrepresent a promising antineoplastic therapy. However, current ADCs face critical challenges, including non-responsive cancers, drug resistance, and rapid patient relapse, primarily driven by tumor heterogeneity and resistance. To address these issues, dual-payload ADCs have emerged as an innovative strategy: they deliver two cytotoxic agents to enhance efficacy via synergistic effects, mitigate resistance, and enable flexible dosing. Despite their potential, dual-payload ADC development remains complex; cell panel-based studies and drug-resistant cell lines serve as powerful tools to explore effective dual-payload ADCs by evaluating drug combination synergies and elucidating resistance mechanisms. To advance the development of DDR pathway-based dual-payload ADCs, this study leveraged an in-house DDR platform to conduct large-scale synergy screening in ADC-resistant cell lines. The screening panel included over 120 targeted molecules and chemotherapeutics, covering DDR core pathway proteins (e.g., WRN, PARP, ATR, DNAPK), cell cycle regulators, and kinase families such as tyrosine kinases. Screening results revealed that ATR inhibitors combined with Topoisomerase I (TOPO1) inhibitors exhibit strong synergistic activity; their specific molecular mechanisms require further validation. This optimal combination provides a direct candidate for DDR pathway-targeted dual-payload ADC development, with the potential to accelerate the research, development, and translation of next-generation DDR-enabled ADC modalities.

Signal Transducer and Activator of Transcription 6 (STAT6) is a central mediator of IL‑4/IL‑13 signaling, and its dysregulation contributes to asthma, atopic dermatitis, fibrosis, and other immune‑driven diseases. With STAT6 therapeutics such as KT‑621 and REX‑8756 advancing clinically, there is growing demand for platforms that generate precise, translationally relevant data.
ICE Biosciences has developed a comprehensive, end‑to‑end workflow for STAT6 inhibitor and PROTAC discovery, built on our integrated target- ed protein degradation (TPD) platform and fully compatible with high‑throughput screening (HTS). Orthogonal biochemical and biophysical assays, including dual‑injection SPR for binding‑site resolution, establish robust target engagement. Cellular assays extend this to degradation, employing WB, FACS, and HiBiT readouts. To enhance translational relevance, functional assays using human primary samples, such as cytokine measurements in blood and smooth muscle cells, are developed. Selectivity and cross‑species profiling ensure confidence across models, while in‑depth MoA studies provide mechanistic clarity. Beyond molecular and cellular characterization, the workflow integrates comprehensive safety evaluation with advancing in studies. By uniting biochemical precision, cellular function, and pharmacological depth, this platform delivers actionable insights to accelerate next‑generation STAT6 therapeutics for autoimmune and inflammatory diseases.
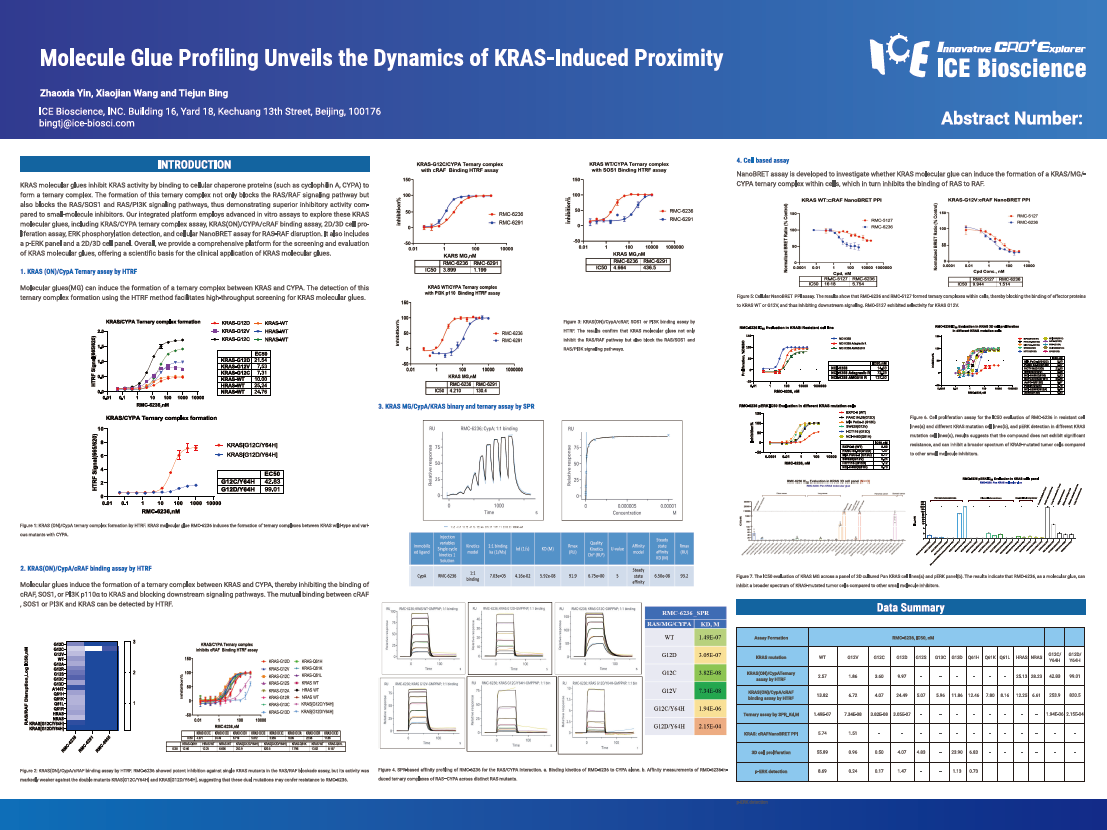
KRAS molecular glues inhibit KRAS activity by binding to cellular chaperone proteins (such as cyclophilin A, CYPA) to form a ternary complex. The formation of this ternary complex not only blocks the RAS/RAF signaling pathway but also blocks the RAS/SOS1 and RAS/PI3K signaling pathways, thus demonstrating superior inhibitory activity compared to small-molecule inhibitors. Our integrated platform employs advanced in vitro assays to explore these KRAS molecular glues, including KRAS/CYPA ternary complex assay, KRAS(ON)/CYPA/cRAF binding assay, 2D/3D cell proliferation assay, ERK phosphorylation detection, and cellular NanoBRET assay for RAS-RAF disruption. It also includes a p-ERK panel and a 2D/3D cell panel. Overall, we provide a comprehensive platform for the screening and evaluation of KRAS molecular glues, offering a scientific basis for the clinical application of KRAS molecular glues.
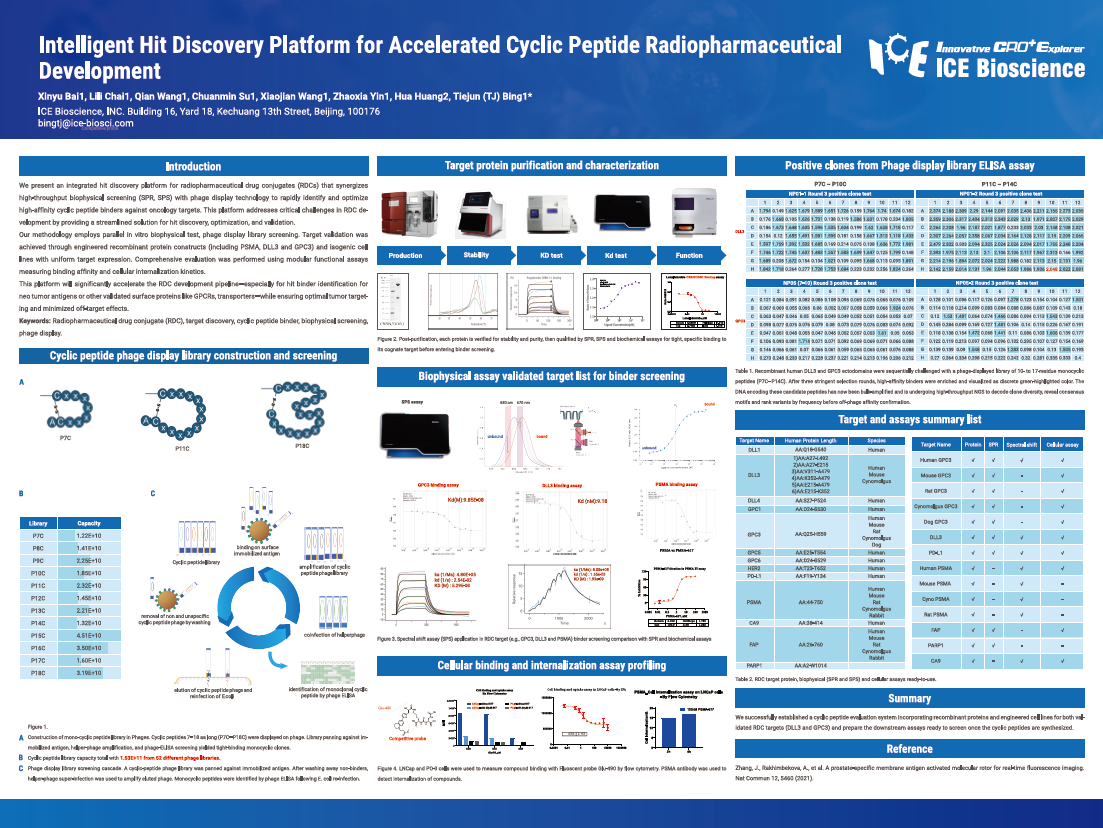
We present an integrated hit discovery platform for radiopharmaceutical drug conjugates (RDCs) that synergizes high-throughput biophysical screening (SPR, SPS) with phage display technology to rapidly identify and optimize high-affinity cyclic peptide binders against oncology targets. This platform addresses critical challenges in RDC development by providing a streamlined solution for hit discovery, optimization, and validation.
Our methodology employs parallel in vitro biophysical test, phage display library screening. Target validation was achieved through engineered recombinant protein constructs (including PSMA, DLL3 and GPC3) and isogenic cell lines with uniform target expression. Comprehensive evaluation was performed using modular functional assays measuring binding affinity and cellular internalization kinetics.
This platform will significantly accelerate the RDC development pipeline—especially for hit binder identification for neo tumor antigens or other validated surface proteins like GPCRs, transporters—while ensuring optimal tumor targeting and minimized off-target effects.