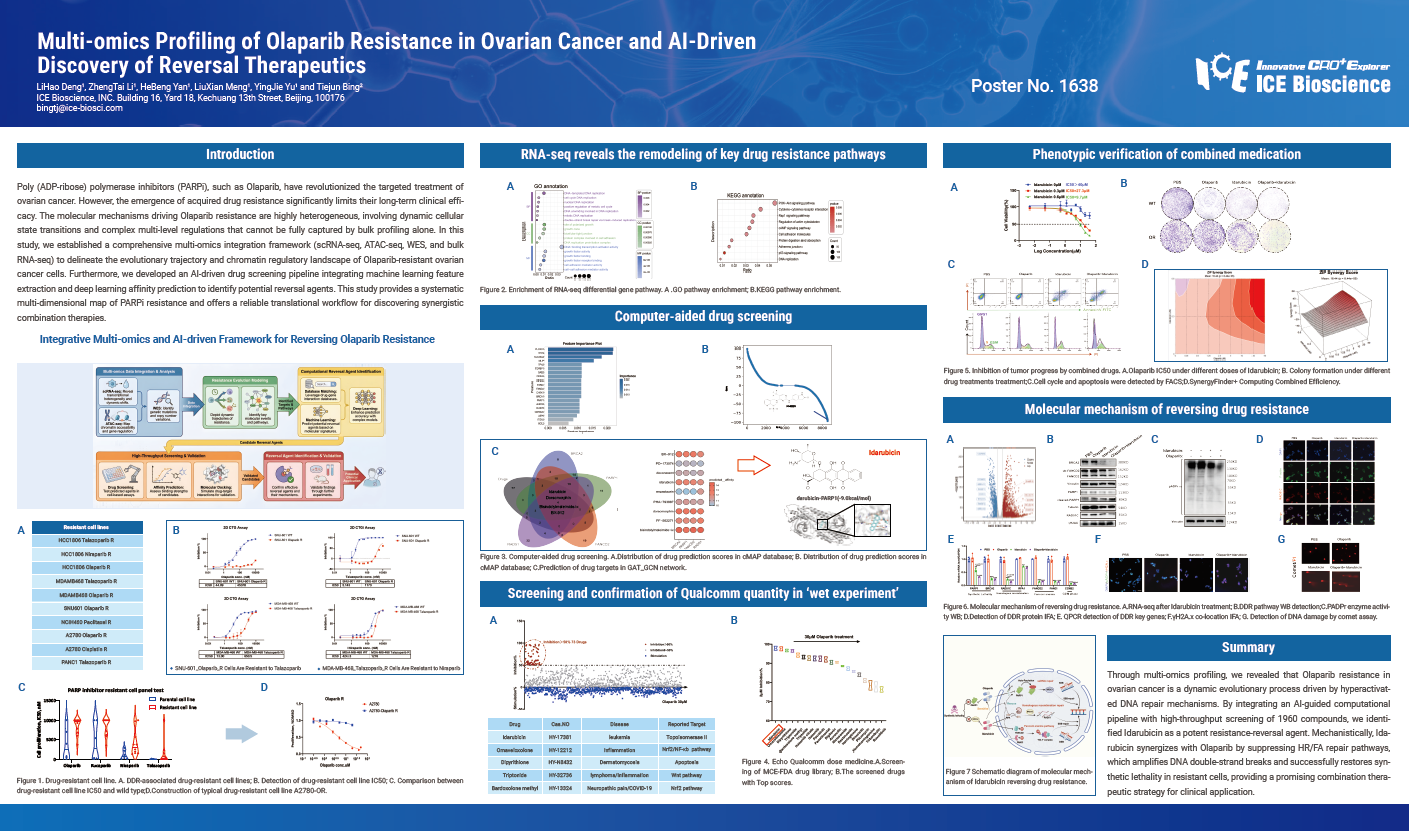
Poly (ADP-ribose) polymerase inhibitors (PARPi), such as Olaparib, have revolutionized the targeted treatment of ovarian cancer. However, the emergence of acquired drug resistance significantly limits their long-term clinical efficacy. The molecular mechanisms driving Olaparib resistance are highly heterogeneous, involving dynamic cellular state transitions and complex multi-level regulations that cannot be fully captured by bulk profiling alone. In this study, we established a comprehensive multi-omics integration framework (scRNA-seq, ATAC-seq, WES, and bulk RNA-seq) to delineate the evolutionary trajectory and chromatin regulatory landscape of Olaparib-resistant ovarian cancer cells. Furthermore, we developed an AI-driven drug screening pipeline integrating machine learning feature extraction and deep learning affinity prediction to identify potential reversal agents. This study provides a systematic multi-dimensional map of PARPi resistance and offers a reliable translational workflow for discovering synergistic combination therapies.
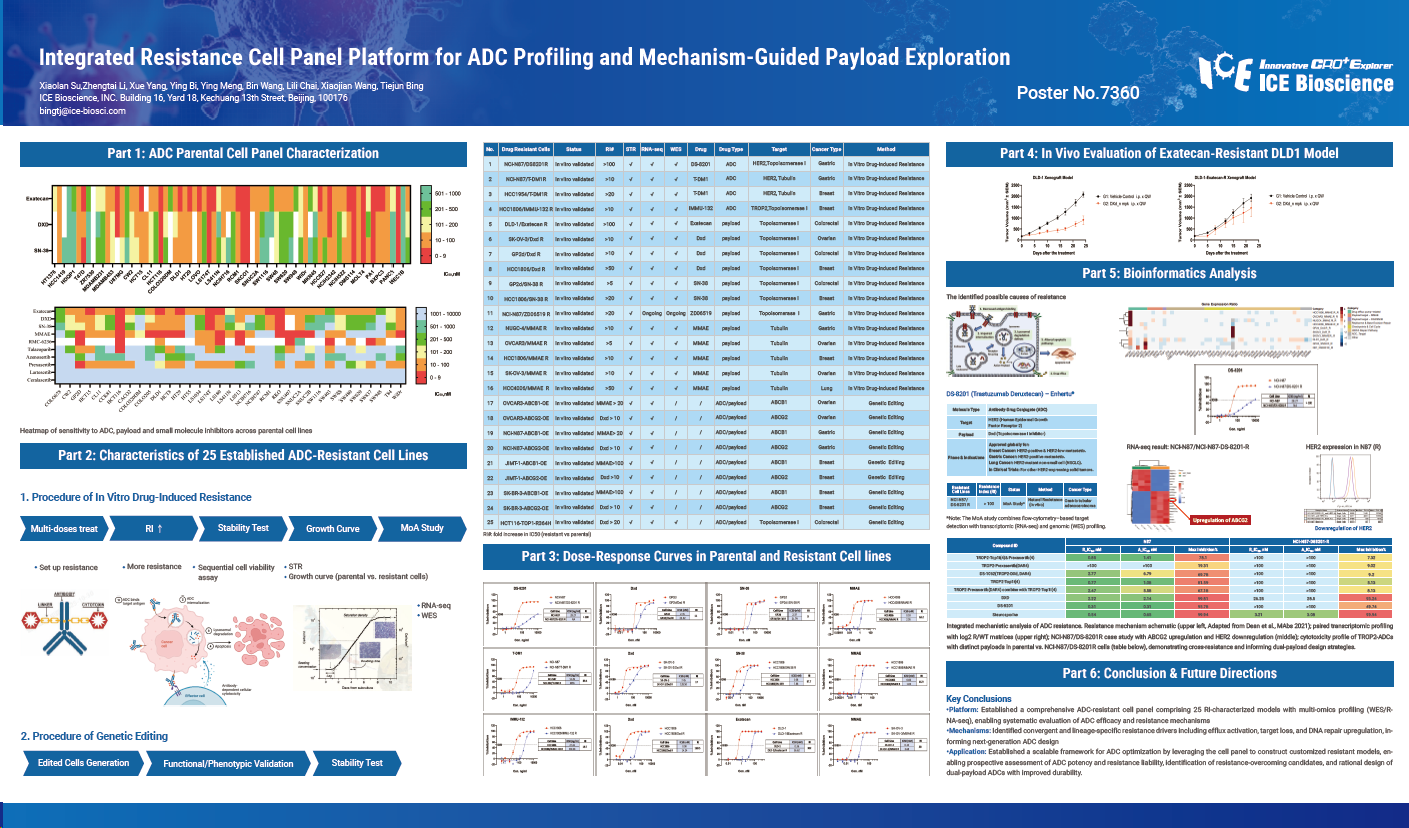
Platform: Established a comprehensive ADC-resistant cell panel comprising 25 RI-characterized models with multi-omics profiling (WES/RNA- seq), enabling systematic evaluation of ADC efficacy and resistance mechanisms.
Mechanisms: Identified convergent and lineage-specific resistance drivers including efflux activation, target loss, and DNA repair upregulation, informing next-generation ADC design.
Application: Established a scalable framework for ADC optimization by leveraging the cell panel to construct customized resistant models, enabling prospective assessment of ADC potency and resistance liability, identification of resistance-overcoming candidates, and rational design of dual-payload ADCs with improved durability.
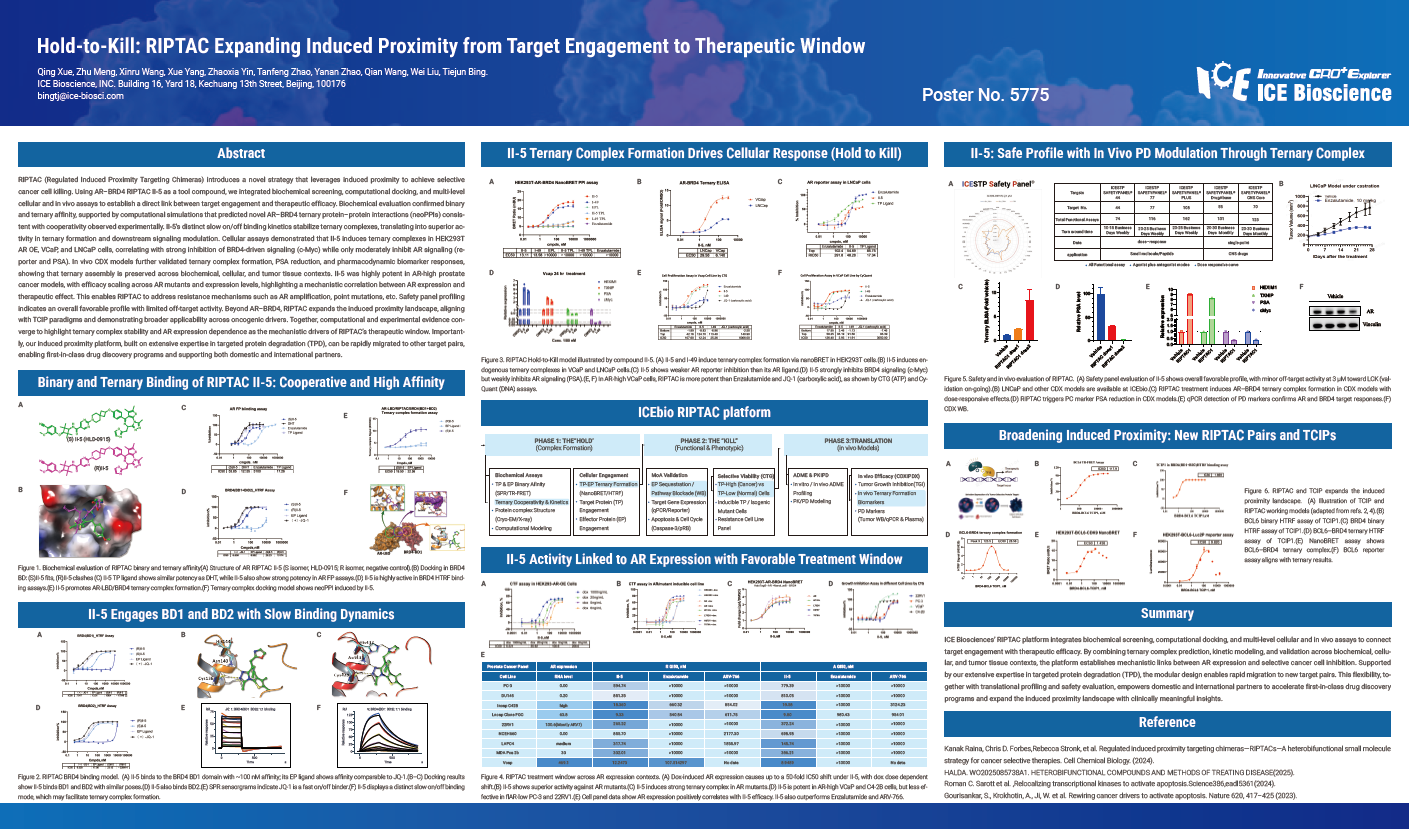
RIPTAC (Regulated Induced Proximity Targeting Chimeras) introduces a novel strategy that leverages induced proximity to achieve selective cancer cell killing. Using AR–BRD4 RIPTAC II-5 as a tool compound, we integrated biochemical screening, computational docking, and multi-level cellular and in vivo assays to establish a direct link between target engagement and therapeutic efficacy. Biochemical evaluation confirmed binary and ternary affinity, supported by computational simulations that predicted novel AR–BRD4 ternary protein–protein interactions (neoPPIs) consistent with cooperativity observed experimentally. II-5’s distinct slow on/off binding kinetics stabilize ternary complexes, translating into superior activity in ternary formation and downstream signaling modulation. Cellular assays demonstrated that II-5 induces ternary complexes in HEK293T AR OE, VCaP, and LNCaP cells, correlating with strong inhibition of BRD4-driven signaling (c-Myc) while only moderately inhibit AR signaling (reporter and PSA). In vivo CDX models further validated ternary complex formation, PSA reduction, and pharmacodynamic biomarker responses, showing that ternary assembly is preserved across biochemical, cellular, and tumor tissue contexts. II-5 was highly potent in AR-high prostate cancer models, with efficacy scaling across AR mutants and expression levels, highlighting a mechanistic correlation between AR expression and therapeutic effect. This enables RIPTAC to address resistance mechanisms such as AR amplification, point mutations, etc. Safety panel profiling indicates an overall favorable profile with limited off-target activity. Beyond AR–BRD4, RIPTAC expands the induced proximity landscape, aligning with TCIP paradigms and demonstrating broader applicability across oncogenic drivers. Together, computational and experimental evidence converge to highlight ternary complex stability and AR expression dependence as the mechanistic drivers of RIPTAC’s therapeutic window. Importantly, our induced proximity platform, built on extensive expertise in targeted protein degradation (TPD), can be rapidly migrated to other target pairs, enabling first-in-class drug discovery programs and supporting both domestic and international partners.
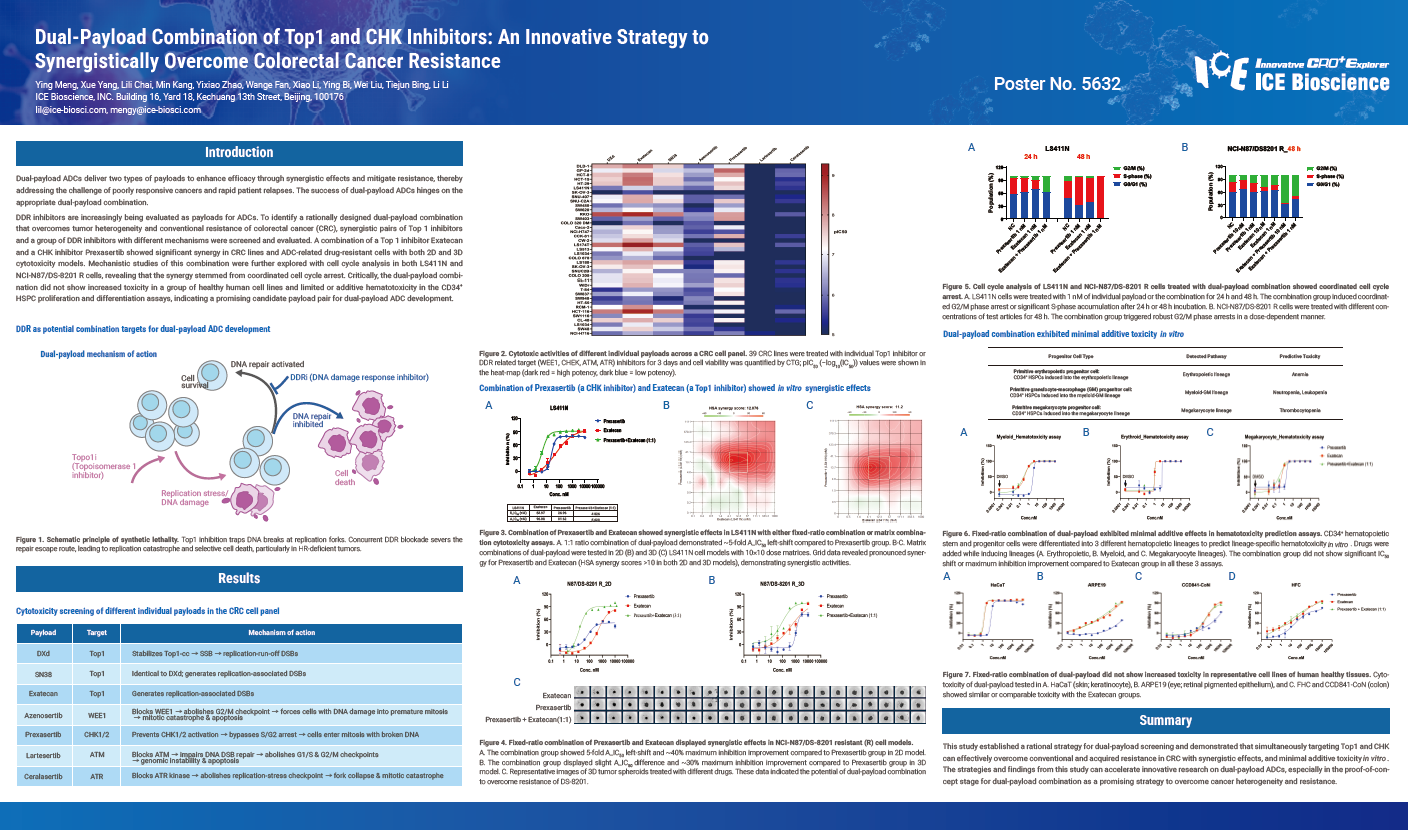
Dual-payload ADCs deliver two types of payloads to enhance efficacy through synergistic effects and mitigate resistance, thereby addressing the challenge of poorly responsive cancers and rapid patient relapses. The success of dual-payload ADCs hinges on the appropriate dual-payload combination.
DDR inhibitors are increasingly being evaluated as payloads for ADCs. To identify a rationally designed dual-payload combination that overcomes tumor heterogeneity and conventional resistance of colorectal cancer (CRC), synergistic pairs of Top 1 inhibitors and a group of DDR inhibitors with different mechanisms were screened and evaluated. A combination of a Top 1 inhibitor Exatecan and a CHK inhibitor Prexasertib showed significant synergy in CRC lines and ADC-related drug-resistant cells with both 2D and 3D cytotoxicity models. Mechanistic studies of this combination were further explored with cell cycle analysis in both LS411N and NCI-N87/DS-8201 R cells, revealing that the synergy stemmed from coordinated cell cycle arrest. Critically, the dual-payload combination did not show increased toxicity in a group of healthy human cell lines and limited or additive hematotoxicity in the CD34+ HSPC proliferation and differentiation assays, indicating a promising candidate payload pair for dual-payload ADC development.