
Induced proximity, which utilizes small molecules to facilitate an interaction between two proteins to harness natural biological pathways, represents a revolution in synthetic chemistry and drug discovery [1]. This area has gained significant clinical traction, with targeted protein degradation leading the way. One example is Molecular glue degraders (MGDs), which induce the proximity of target protein to E3 ubiquitin ligases, leading to target ubiquitination and degradation. The mechanism is different from traditional inhibitors in drug discovery and broaden the approach, especially for the target used to be considered "undruggable". MRT-6160, a first-in-class VAV1-directed MGDs currently in clinical phase I, showed the remarkable potential in Immunology and inflammatory diseases such as rheumatoid arthritis and colitis [2][3]. Current advancements highlight its ability to reduce proinflammatory cytokine production, inhibit pathogenic T cell polarization, and mitigate autoimmune responses.

In the development of new drugs, it is crucial to know the safety profile of drug candidates in advance, as this can prevent the drug from being withdrawn from the market due to safety concerns after launch, thus reducing the risk for pharmaceutical companies.
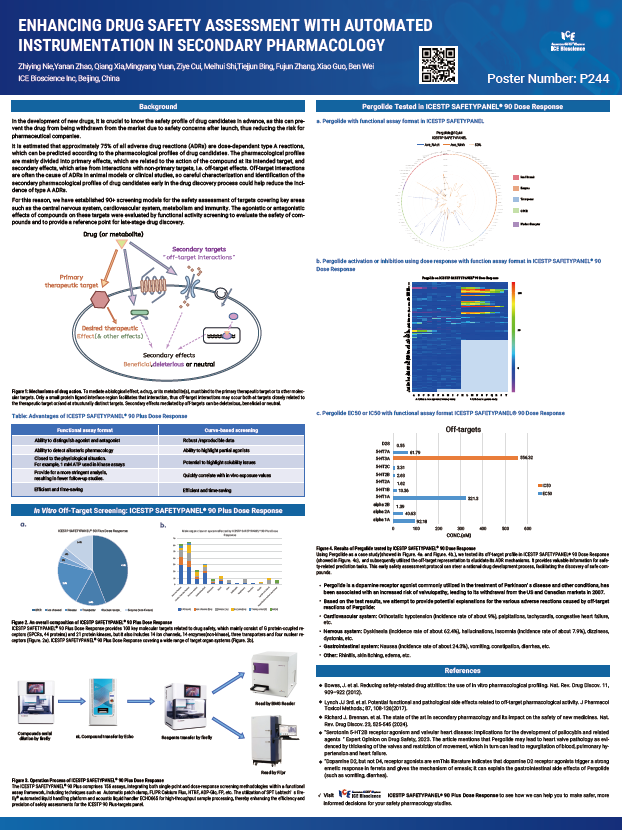
It is estimated that approximately 75% of all adverse drug reactions (ADRs) are dose-dependent type A reactions, which can be predicted according to the pharmacological profiles of drug candidates. The pharmacological profiles are mainly divided into primary effects, which are related to the action of the compound at its intended target, and secondary effects, which arise from interactions with non-primary targets, i.e. off-target effects. Off-target interactions are often the cause of ADRs in animal models or clinical studies, so careful characterization and identification of the secondary pharmacological profiles of drug candidates early in the drug discovery process could help reduce the incidence of type A ADRs.

High-throughput screening of a cancer cell panel represents a pivotal instrument in the armamentarium of drug discovery and development. The sensitivity profiles of these cell lines serve as a cornerstone for the refinement of in vivo model systems and for the stratification of patient cohorts in clinical trials. Werner Syndrome RecQ Helicase (WRN), a member of the RecQ helicase family, is instrumental in the preservation of genomic integrity. Its multifaceted role encompasses DNA repair, replication, recombination, and telomere maintenance. WRN has garnered significant attention as a potential therapeutic nexus in oncology, particularly within the context of tumors exhibiting microsatellite instability (MSI). The synthetic lethality between WRN and MSI has been the subject of intensive research, demonstrating that WRN inhibition is selectively toxic to cancer cells harboring a high microsatellite instability (MSI-H) phenotype, which is a characteristic observed across a spectrum of malignancies, including colorectal, endometrial, and gastric cancers.
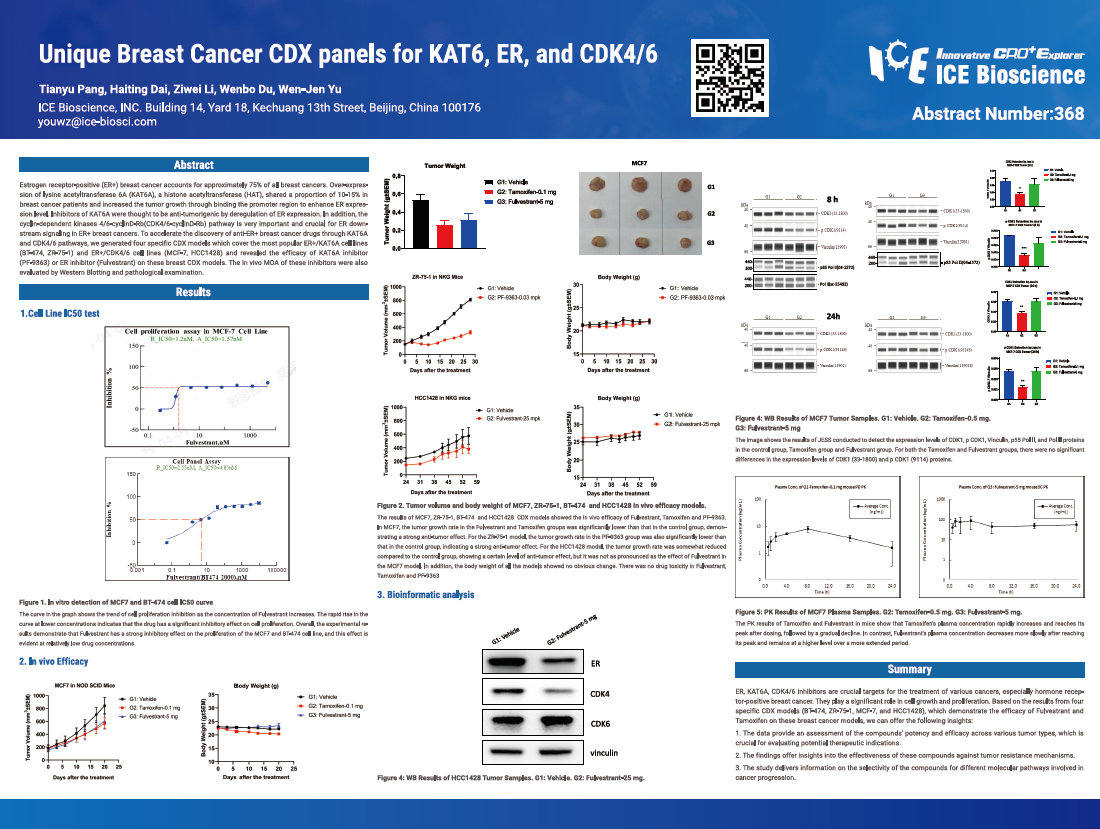
Estrogen receptor-positive (ER+) breast cancer accounts for approximately 75% of all breast cancers. Over-expression of lysine acetyltransferase 6A (KAT6A), a histone acetyltransferase (HAT), shared a proportion of 10-15% in breast cancer patients and increased the tumor growth through binding the promoter region to enhance ER expression level. Inhibitors of KAT6A were thought to be anti-tumorigenic by deregulation of ER expression. In addition, the cyclin-dependent kinases 4/6-cyclinD-Rb (CDK4/6-cyclinD-Rb) pathway is very important and crucial for ER downstream signaling in ER+ breast cancers. To accelerate the discovery of anti-ER+ breast cancer drugs through KAT6A and CDK4/6 pathways, we generated four specific CDX models which cover the most popular ER+/KAT6A cell lines (BT-474, ZR-75-1) and ER+/CDK4/6 cell lines (MCF-7, HCC1428) and revealed the efficacy of KAT6A inhibitor (PF-9363) or ER inhibitor (Fulvestrant) on these breast CDX models. The in vivo MOA of these inhibitors were also evaluated by Western Blotting and pathological examination.