Characterizing a RIPTAC requires more than confirming that a bifunctional molecule binds within the system. A meaningful biochemical workflow should determine whether each binding element remains active after linker incorporation, whether that activity is preserved across relevant target formats, whether the molecule supports ternary assembly rather than independent binding alone, and whether the resulting complex is associated with a measurable target-proximal biochemical consequence.
Publicly, HLD-0915 has advanced into Phase 1/2 clinical development for metastatic castration-resistant prostate cancer (mCRPC), with first patient dosing reported in 2025, FDA Fast Track designation granted later that year, and initial first-in-human data subsequently disclosed. Following Halda’s acquisition by Johnson & Johnson in late 2025, the program became part of J&J’s oncology pipeline.
For scientists working on induced proximity programs, this case illustrates how ICE Bioscience can support custom biochemical assay development around program-specific mechanistic questions.
For BRD4, binary binding was intentionally resolved across BD1, BD2, and BD1+BD2 because BRD4 contains two bromodomains with related but non-identical ligand recognition properties. In bifunctional molecule design, linker incorporation can shift the binding profile across these domains, so a single aggregated BRD4 readout may miss mechanistically relevant differences. This level of domain resolution is therefore target- and question-dependent rather than a universal requirement for every epigenetic protein program.
In this case, separating BD1 and BD2 provides a clearer view of how II-5 behaves across distinct BRD4 binding formats. The combined BD1+BD2 construct adds further context by showing compound behavior when both domains are presented within the same target format.
The comparison between (S)II-5 and (R)II-5 is especially informative. In RIPTAC characterization, a matched stereochemical pair helps test whether the observed profile depends on precise three-dimensional presentation rather than broad nonspecific interaction. Here, the active stereoisomer shows measurable binding, while the opposite stereoisomer loses BRD4 activity, which is consistent with a structure-dependent mechanism.
AR engagement was assessed using a fluorescence polarization assay selected as a suitable biochemical format for AR-LBD. The workflow therefore applies fit-for-purpose biochemical formats to distinct mechanistic questions across the characterization cascade.
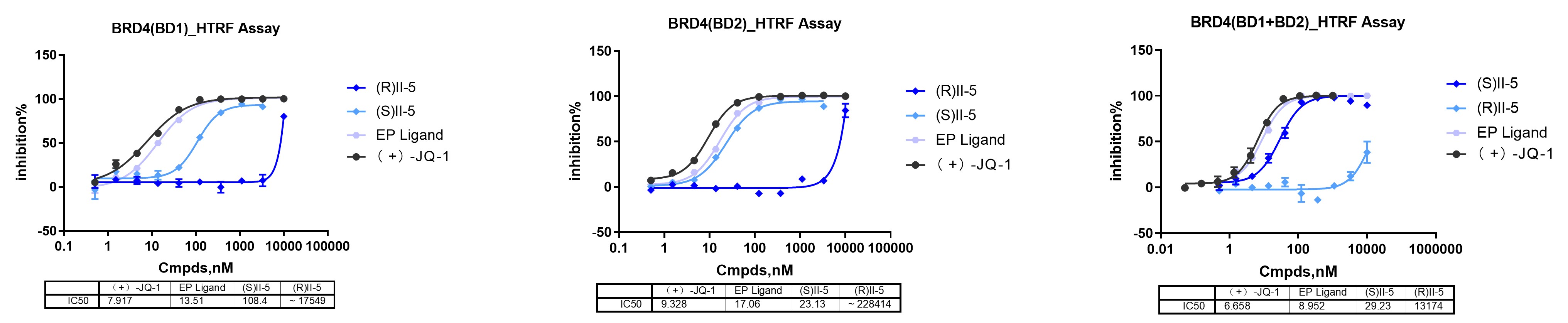
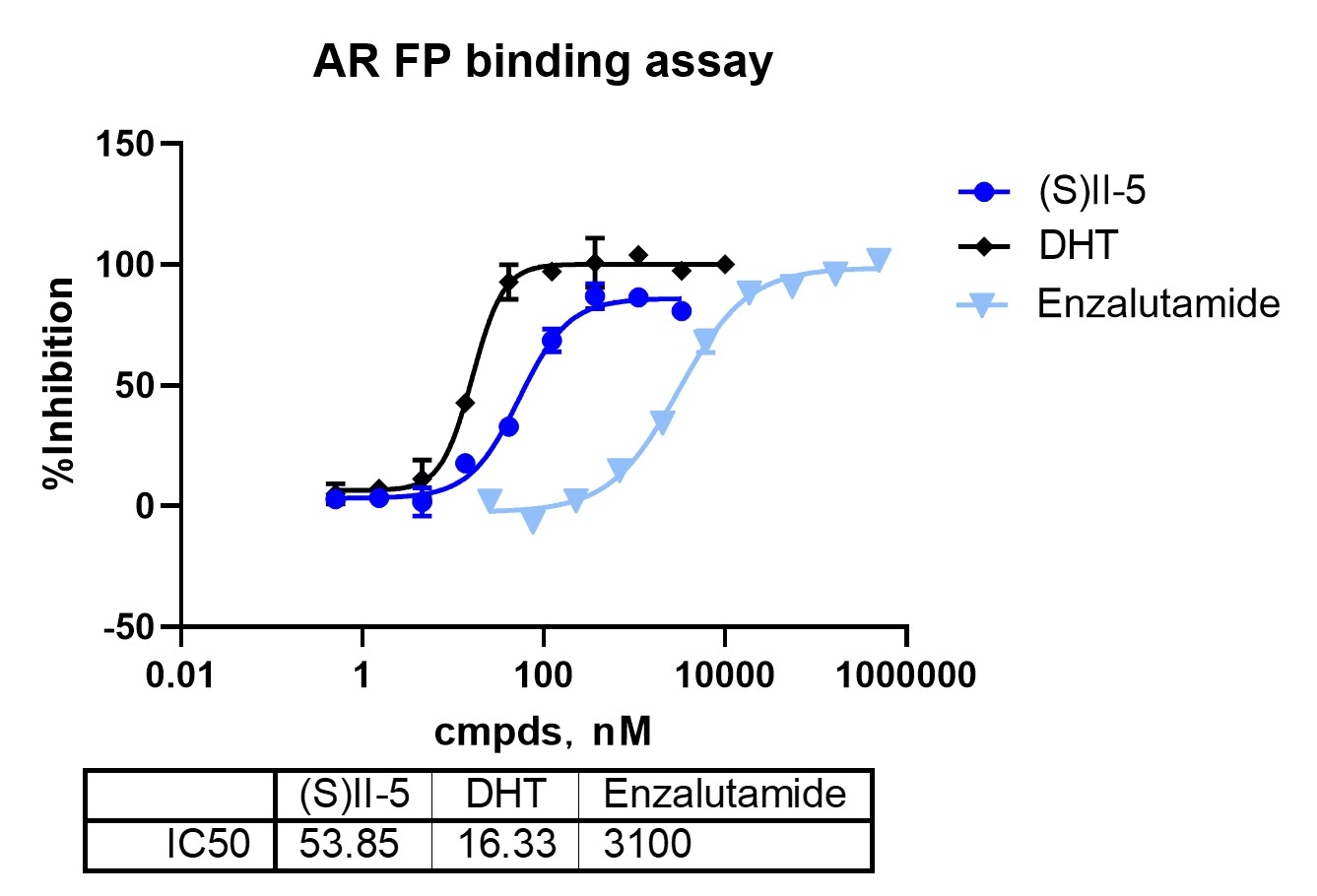
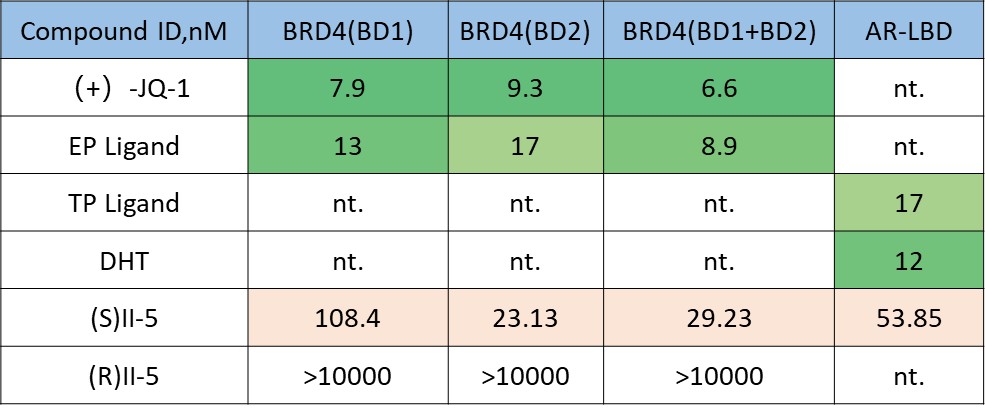
Binary binding assays establish whether both recognition elements remain functional after bifunctional assembly. In this study, II-5 retained measurable interaction with BRD4 and AR-LBD, while showing a differentiated BRD4 domain profile across BD1, BD2, and BD1+BD2. The stereochemical control further supports a structure-dependent signal.
Binary binding alone, however, does not establish that a RIPTAC is mechanistically effective. Independent engagement of two targets does not guarantee that the molecule can position them in a way that supports induced proximity. For that reason, the binary layer is a necessary entry point, but it is not a sufficient basis for advancement.
The central question in RIPTAC characterization is whether a bifunctional molecule can do more than bind two proteins independently. The ternary assay addresses that question more directly by testing for a ternary assembly signal consistent with AR–RIPTAC–BRD4 complex formation under the assay conditions.
This step separates outcomes that may otherwise look similar in early screening: compounds that bind one target, compounds that bind both targets, and compounds that also support induced proximity. Structural changes that preserve binary binding may still weaken ternary assembly, while molecules with only moderate binary potency may remain highly relevant when their geometry supports productive spatial organization.
In this dataset, the comparison between active and inactive stereochemical variants is especially informative. It supports the view that ternary behavior depends on molecular architecture rather than simple target occupancy alone.
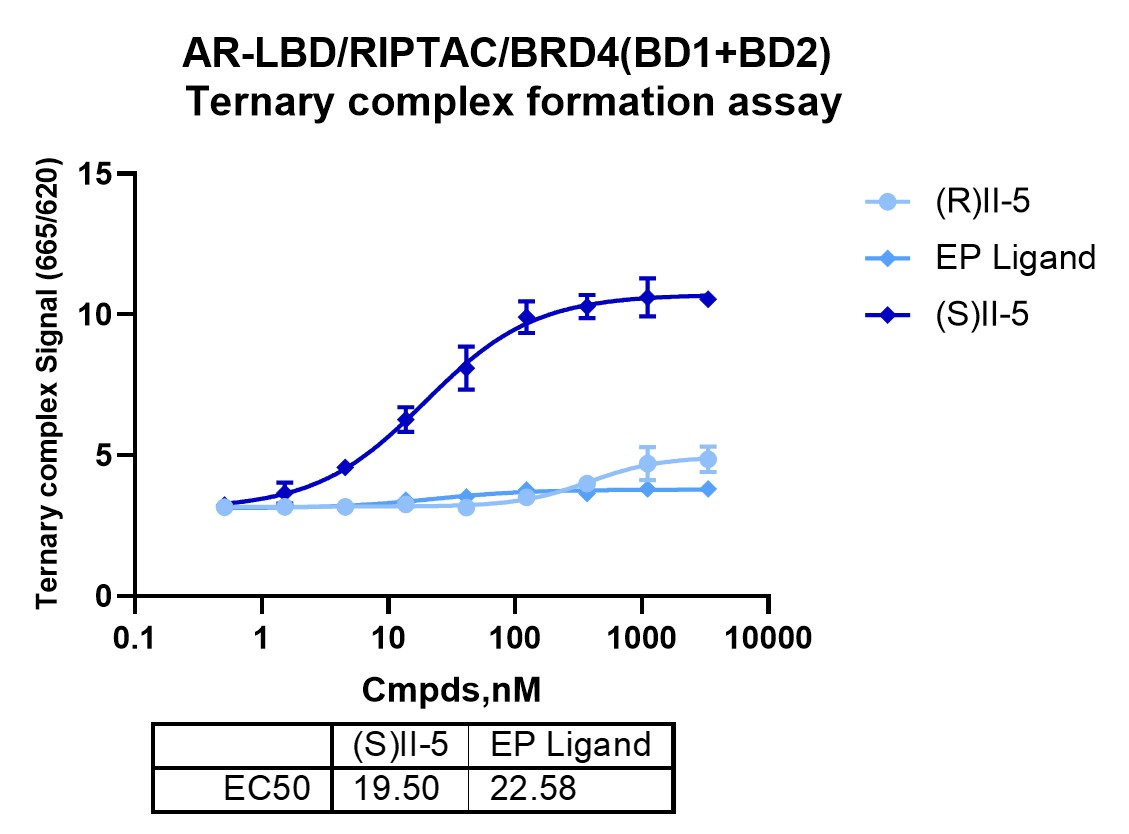
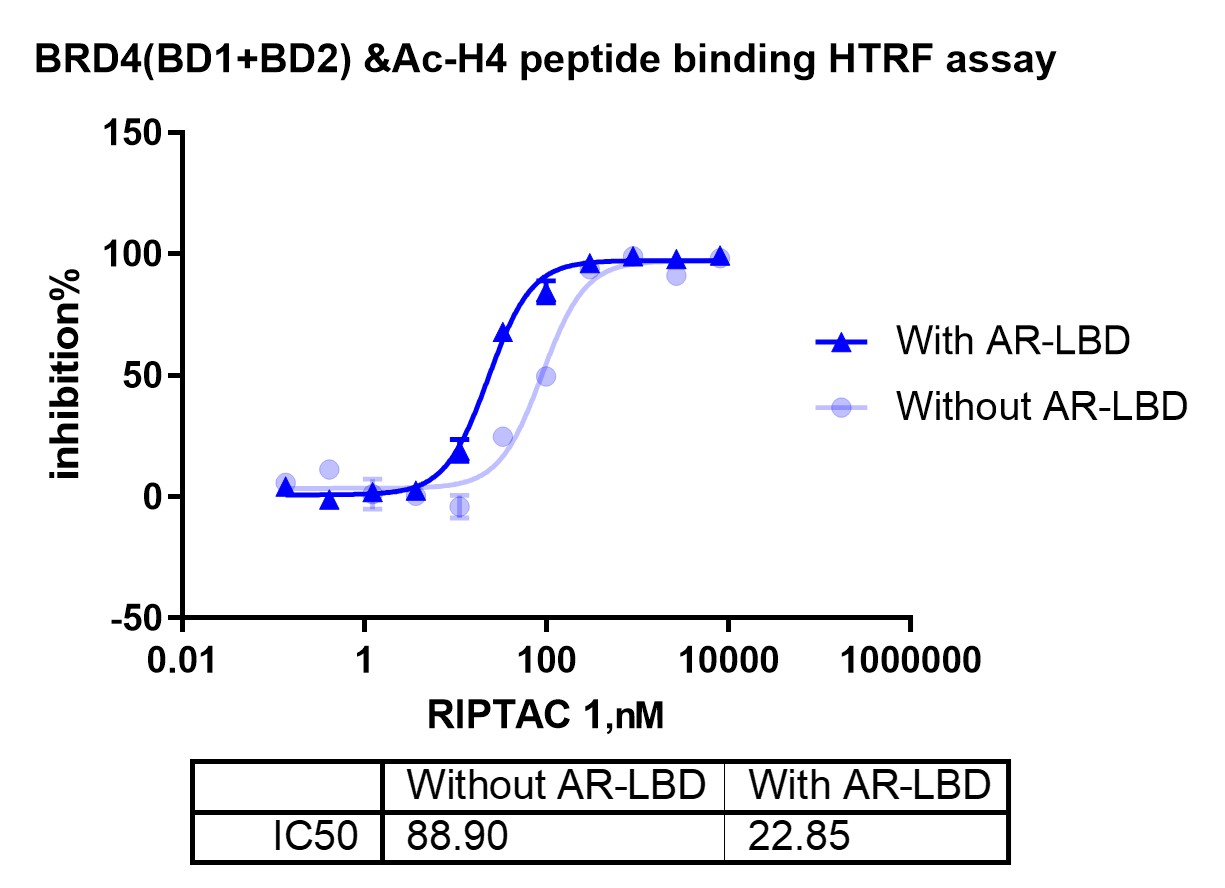
Ternary formation is a stronger mechanistic filter than binary binding, but it does not by itself show whether BRD4 engagement perturbs a functionally relevant BRD4 recognition event. In this workflow, that additional layer was addressed through an Ac-H4 peptide binding assay, which measures the ability of BRD4 to interact with an acetylated histone H4 peptide substrate mimic.
This assay is functionally relevant because the bromodomains of BRD4 recognize acetyl-lysine marks on histone tails. An acetylated H4 peptide therefore serves as a biochemical proxy for a native BRD4 recognition event rather than a generic tracer. When a BRD4-binding element occupies the bromodomain in a relevant manner, it can reduce BRD4 association with the Ac-H4 peptide. In that sense, the readout reflects a target-proximal biochemical consequence of BRD4 engagement.
The value of this step lies in narrowing the gap between ternary assembly and later cellular evaluation. When binary binding, ternary formation, and Ac-H4 peptide displacement align, the dataset provides stronger support for prioritizing a molecule with both ternary competence and a target-proximal biochemical consequence on the BRD4 side of the system.
In this AR–BRD4 RIPTAC case study, BRD4 domain-resolved binding, AR engagement, ternary complex assessment, and biochemical functional testing were arranged as a sequence of increasingly discriminating readouts. This design provides a clearer basis for distinguishing simple target engagement from ternary competence and target-proximal biochemical consequence, and therefore a better basis for RIPTAC prioritization than binary binding alone.
2026-05-13
2025-10-30
2025-10-23
2025-09-28
We value your inquiries and are here to provide you with tailored solutions for your drug discovery and development needs. Whether you have questions, require more information, or are interested in discussing potential collaborations, our team of experts is just a message away.
Feel free to reach out to us.